TUNA: Multimodal Target-aware Network for Protein-Ligand Affinity Prediction
More Links
Why Binding Affinity Prediction Still Needs Better Context
Predicting how strongly a drug molecule binds to its target protein is a central challenge in early-stage drug discovery. Traditional structure-based models can be highly accurate, but they rely on experimentally determined 3D protein-ligand complexes, which are unavailable for most proteins. Sequence-based deep learning models scale better, but they often miss a critical piece of biological context: the local binding pocket where the interaction actually occurs. The paper “TUNA: A Target-aware Unified Network for Protein-Ligand Binding Affinity Prediction via Multi-Modal Feature Integration” introduces TUNA, a model designed to bridge this gap by combining the scalability of sequence-based methods with pocket-level target awareness
Integrating Proteins, Pockets, and Ligands in One Model
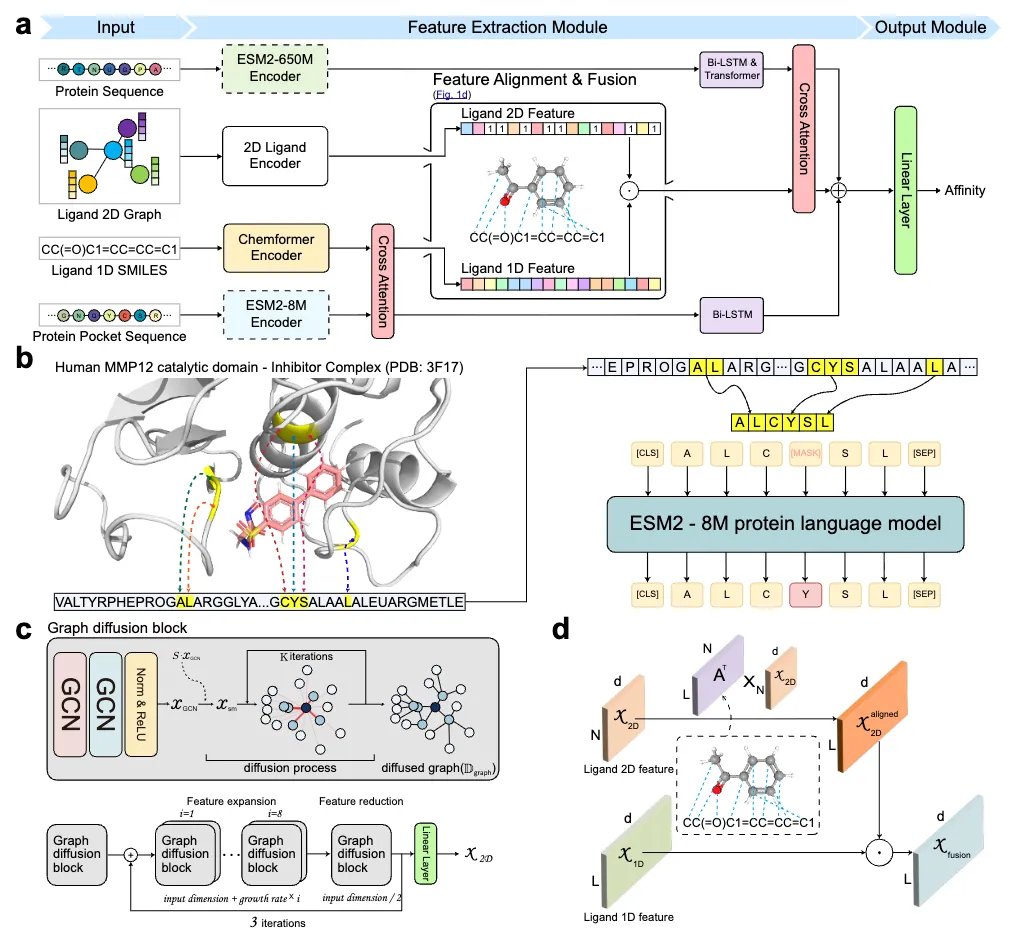
The core idea behind TUNA is multi-modal integration. On the protein side, the model uses pretrained protein language models (ESM2) to encode both the full protein sequence and localized pocket sequences. When experimental pocket annotations are unavailable, TUNA infers protein structures using AlphaFold and detects candidate pockets computationally. On the ligand side, TUNA jointly models SMILES strings, which capture chemical syntax, and molecular graphs, which encode structural connectivity. A dedicated alignment and fusion strategy ensures that symbolic (1D) and structural (2D) ligand features are combined in a consistent and biologically meaningful way.
Cross-Attention for Accuracy and Interpretability
A key strength of TUNA lies in how these modalities interact. The model uses cross-attention mechanisms that allow ligand features to attend to protein pocket representations and then refine global protein features based on ligand context. This design improves predictive accuracy while also enhancing interpretability. By analyzing attention weights, TUNA can highlight protein residues that are likely involved in binding, providing insights into potential binding sites rather than producing only a single affinity score.
For detailed methodology and findings, read our full paper here.