SpaceExpress: A Statistical Framework for Comparative Spatial Transcriptomics
More Links & Resources
Unlocking the Secrets of Spatial Gene Expression with SpaceExpress
Understanding how genes express themselves spatially across different tissues is essential for grasping how biology works at a cellular level. But there’s a challenge: comparing spatial gene expression across different conditions (like disease vs. healthy tissue, or different behavioral states) is notoriously tricky because tissues naturally vary in structure and orientation. That’s where our new tool, SpaceExpress, steps in.
What is SpaceExpress?
SpaceExpress is a computational tool we developed to analyze spatial transcriptomics data (a type of data that shows not only which genes are active but also where exactly they’re active within tissues). Our method cleverly embeds cells from different tissue samples into a shared coordinate system, allowing for meaningful comparisons even when samples differ structurally.
Instead of traditional coordinates (x, y, z), SpaceExpress creates intrinsic “biological coordinates” based on gene expression patterns. This makes comparisons robust and biologically relevant, highlighting precisely which genes change their spatial expression patterns between conditions.
How does SpaceExpress work?
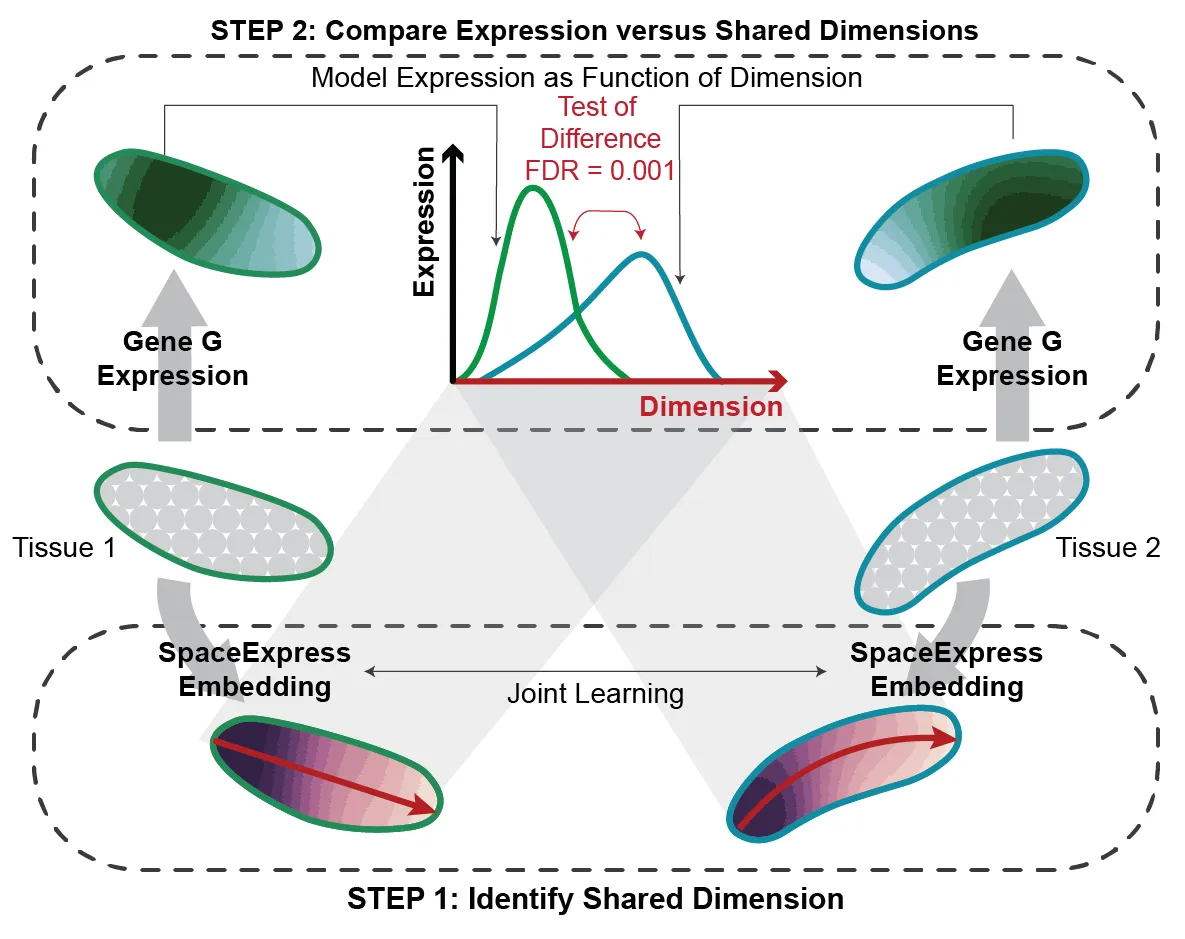
- Step 1: Learning biological coordinates: Using a neural network, it translates this spatial structure into a set of latent dimensions—think of them as “biological axes”—that consistently describe tissue organization across different samples.
- Step 2: Identifying Differential Spatial Expression (DSE): Once the shared biological coordinates are set, it uses advanced statistical methods to pinpoint genes whose spatial patterns significantly differ across conditions.
Applications & Discoveries
We validated SpaceExpress with both synthetic data and real biological examples, including datasets from:
- Honey Bee Brains: By comparing soldier and forager bees, we identified specific genes with spatial patterns connected to behavioral differences, revealing previously unseen spatial neurogenomic signatures linked to complex behaviors.
- Mouse Brains: Analyzing regions associated with aggressive behaviors, SpaceExpress found notable genes like oxytocin and tyrosine hydroxylase whose spatial expression patterns differ significantly based on behavioral conditions.
These insights offer potential pathways to understanding how spatial gene expression impacts behaviors and diseases.
Why This Matters
Spatial transcriptomics is reshaping biology by connecting gene activity with physical tissue structure. SpaceExpress bridges crucial gaps in current analytical methods, helping scientists rigorously pinpoint which spatial gene expression changes matter most for biology and medicine.
Our tool doesn’t just answer technical questions, it opens new doors to exploring how our biology is wired at the deepest levels, enabling discoveries that were previously impossible.
Explore our paper to dive deeper into the revolutionary capabilities of SpaceExpress!