MANTIS: Analytics toolkit for spatial metabolomics with matching spatial transcriptomics data
More Links
Why Spatial Metabolomics Needs Better Analytics
Spatial metabolomics (SM) is rapidly emerging as a powerful technology for mapping metabolic states directly within tissues. Its impact becomes even greater when paired with spatial transcriptomics (ST), which provides complementary information about gene expression, cell types, and tissue organization. However, while several tools exist for aligning and visualizing SM and ST data, there has been limited support for rigorously analyzing biological relationships between metabolites and genes in a spatially aware manner. The paper “MANTIS: Analytics toolkit for spatial metabolomics with matching spatial transcriptomics data” introduces MANTIS, a computational framework designed to fill this gap by focusing on statistically robust discovery of spatial metabolic patterns and gene–metabolite associations.
A Unified Framework for Spatial Patterns and Associations
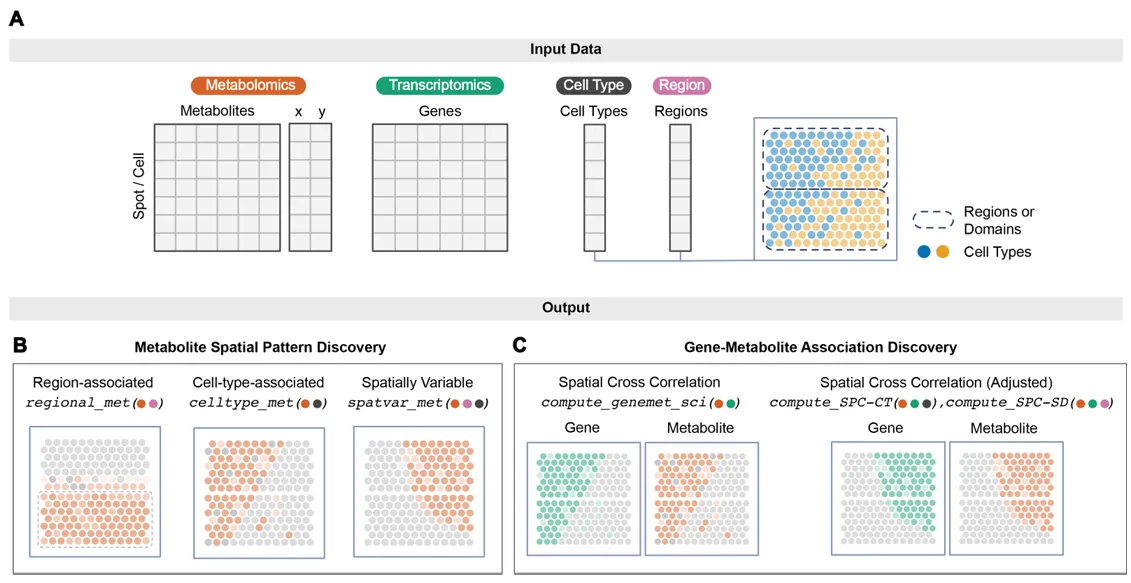
MANTIS is built to analyze paired and aligned SM+ST datasets at the level of cells or spatial spots, optionally incorporating cell type annotations and spatial domains. The toolkit supports multiple types of spatial pattern discovery, including region-associated metabolites, cell type-associated metabolites, and spatially variable metabolites that do not neatly align with known tissue regions or cell types. Importantly, MANTIS explicitly accounts for spatial autocorrelation, a common feature of spatial omics data that can otherwise inflate false positives. By using permutation-based null models that preserve spatial structure, MANTIS provides more conservative and interpretable results than many existing approaches.
Spatially Aware Gene–Metabolite Relationships
One of the most distinctive contributions of MANTIS is its treatment of gene–metabolite associations. Rather than relying on standard correlations such as Pearson’s correlation coefficient, MANTIS uses a spatial cross-correlation statistic to quantify co-variation between genes and metabolites across neighboring spatial locations. The framework further introduces spatial partial correlation to disentangle associations driven purely by shared cell type composition or spatial domains from those that reflect more specific biological relationships. This allows researchers to separate broad co-localization effects from potentially mechanistic gene–metabolite interactions, which is critical for biological interpretation
For detailed methodology and findings, read our full paper here.