MAD: Microenvironment-Aware Distillation for Virtual Spatial Omics from Microscopy
More Links
Why Bridging Microscopy and Omics Still Needs a New Idea
Microscopy and omics each reveal a different side of a cell: imaging captures spatial morphology and tissue context at single-cell resolution, while spatial transcriptomics and related technologies capture molecular states. The dream is to read molecular states directly from routine images without the cost and throughput limits of omics assays. Supervised image-to-omics models have shown this is possible, but they rely on scarce paired datasets and often fail to generalize across tissues, stains, or tasks. The paper “MAD: Microenvironment-Aware Distillation” proposes a self-supervised alternative that lifts the label bottleneck and learns cell-centric representations from unlabeled microscopy at scale.
Dual-View Joint Self-Distillation
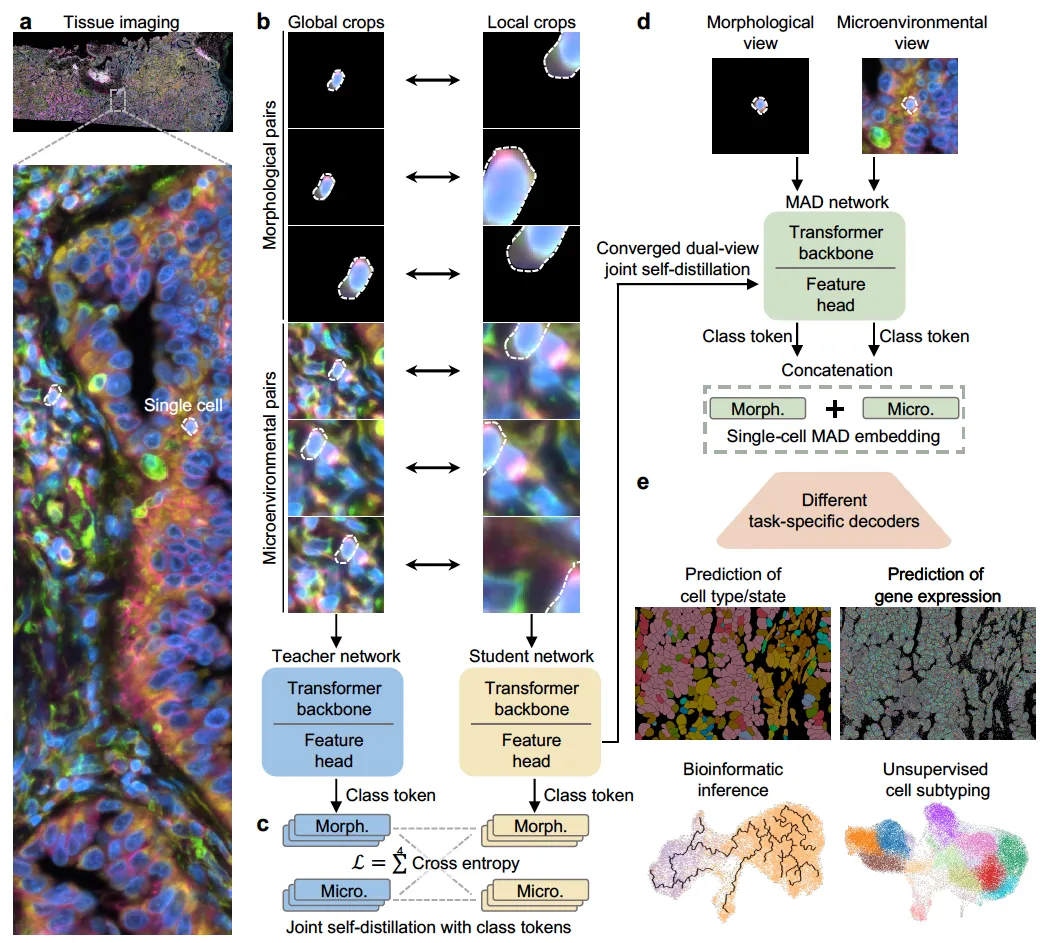
The core idea of MAD is that a cell’s identity is shaped by both intrinsic morphology and its surrounding microenvironment. Prior self-supervised efforts tended to pick one: either single-cell models restricted to simplified culture images, or tissue-level models whose embeddings are limited to the patch or slide level. MAD instead constructs two complementary, cell-centric views for every indexed cell (a morphology view containing only the segmented cell, and a microenvironment view containing the cell within its local neighborhood) and trains a vision transformer to align them in a unified embedding space. Following a DINO-style teacher-student scheme, MAD uses a four-way cross-entropy objective that aligns class tokens across the two views from both networks. The resulting embedding is the concatenation of the morphology and microenvironment class tokens, giving a context-aware, scale-invariant, single-cell representation.
State-of-the-Art on Downstream Tasks with a Label-Efficient Strategy
Across diverse tissues and imaging modalities, MAD achieves state-of-the-art performance on a range of downstream tasks (cell subtyping, transcriptomic prediction, and bioinformatic inference) using lightweight task-specific decoders on top of frozen MAD embeddings. Benchmarked against foundation models with a comparable number of parameters on H&E tissue images, MAD outperforms models that were pretrained on substantially larger datasets, suggesting that the dual-view joint distillation objective captures more biologically relevant structure per unit of data than plain scale. Because the pretraining is self-supervised, MAD directly addresses the label bottleneck that limits supervised image-to-omics methods, offering a practical path toward virtual spatial omics from the microscopy archives that already exist.
For detailed methodology and findings, read our full paper here.